New publication in eLife of the Jordan Lab in collaboration with the IBMB Imaging Platform showing that…
Single-chain dimers from de novo immunoglobulins as robust scaffolds for multiple binding loops
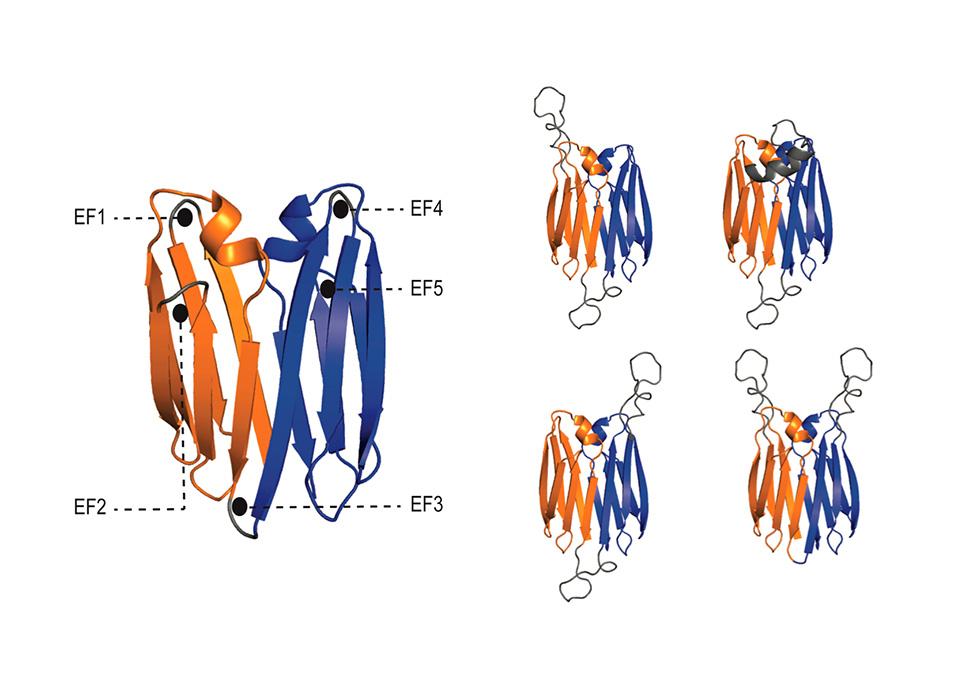
Dr. Enrique Marcos’ laboratory has developed principles for de novo designing single-chain immunoglobulin dimers as new antibody-like format with interfaces diverging from those seen in antibodies, showing enhanced stability, structural accuracy, and both robustness and modularity for harbouring multiple functional loops.
Abstract
Antibody derivatives have sought to recapitulate the antigen binding properties of antibodies, but with improved biophysical attributes convenient for therapeutic, diagnostic and research applications. However, their success has been limited by the naturally occurring structure of the immunoglobulin dimer displaying hypervariable binding loops, which is hard to modify by traditional engineering approaches. Here, we devise geometrical principles for de novo designing single-chain Ig dimers, as a tunable two-domain immunoglobulin architecture that optimizes biophysical properties through more favorable dimer interfaces. Guided by these principles, we computationally designed protein scaffolds that were hyperstable, structurally accurate and robust for accommodating multiple functional loops, both individually and in combination, as confirmed through biochemical assays and X-ray crystallography. We showcase the modularity of this architecture by deep-learning-based diversification, opening up the possibility for tailoring the number, positioning, and relative orientation of ligand-binding loops targeting one or two distal epitopes. Our results provide a route to custom-design robust protein scaffolds for harboring multiple functional loops.
Reference:
Roel-Touris, J., Nadal, M. & Marcos, E. Single-chain dimers from de novo immunoglobulins as robust scaffolds for multiple binding loops. Nat Commun 14, 5939 (2023). https://doi.org/10.1038/s41467-023-41717-5