The Molecular Imaging Platform (MIP) has been set up as a co-operation between the Molecular Biology Institute of Barcelona (IBMB) and the Barcelona Science Park (PCB). We offer a collection of state-of-the-art light microscopy systems and techniques and provide assistance with all aspects of image acquisition and analysis, ranging from advice on imaging experiments to microscopy training or project collaboration.
Available applications
- Automated Wide-Field Microscopy/High content screening
- Live 3D imaging/On-line computational clearing
- Laser Scanning Confocal Microscopy
- Multiphocal High-Speed Confocal Microscopy
- Multiphoton Microscopy/laser cutting/spectral imaging
- Fluorescence Correlation & Cross-correlation Spectroscopy
- Total Internal Reflexion Fluorescence
- Superesolution SRRF-stream
- Image processing and analysis
Modus operandi
- The Platform is accessible 24 hours a day from Monday to Sunday to researchers at IBMB and other PCB and outside institutions and companies.
- We provide System Introductions on a regular basis, where you will be taught in both theoretical and practical aspects of the different image acquisition techniques.
- The equipment can be operated on a self-service mode. Booking permissions are only granted to autonomous users who have been introduced to the systems.
- For further information contact the MIP manager at tgbmco@ibmb.csic.es or mip@ibmb.csic.es
Location
You can find us at the PCB Helix building, base floor PBA07. C/ Baldiri i Reixac 15-20, 08028 Barcelona.
GENERAL EQUIPMENT

CONFOCAL STELLARIS 8 FALCON-FLIM (LEICA MICROSYSTEMS)
The Leica STELLARIS 8 FALCON spectral confocal microscope is equipped with a pulsated white light laser and 5 highly sensitive Power HyD detectors that allows for multiplexing. The FALCON module provides accurate lifetime data for Fluorescence lifetime imaging microscopy (FLIM) and Fluorescence resonance energy transfer (FRET) experiments. The system also comes with the TauSense technology that gives instant and easy access to lifetime-based information. In addition, the system is configured for Fluorescence recovery after photobleaching (FRAP) and for Fluorescence Correlation Spectroscopy (FCS) assays. It is also equipped with an Okolab incubator for live cells experiments.

Ayuda EQC2021-007200-P financiada por:

THUNDER IMAGER 3D LIVE CELL (LEICA MICROSYSTEMS)
Thunder Imager is a new generation widefield technology for fast 3D optical sectioning of fixed and live specimens, from cells up to large tissues, embryos and organoids. Together with a super fast integrated hardware, the Thunder algorithms remove the out-of-focus light in real time, thus opening real new possibilities in high-content applications. Our Thunder system has been appointed as reference machine, the first of its kind in Spain, based on a collaboration agreement with Leica Microsystems.

Specifications:
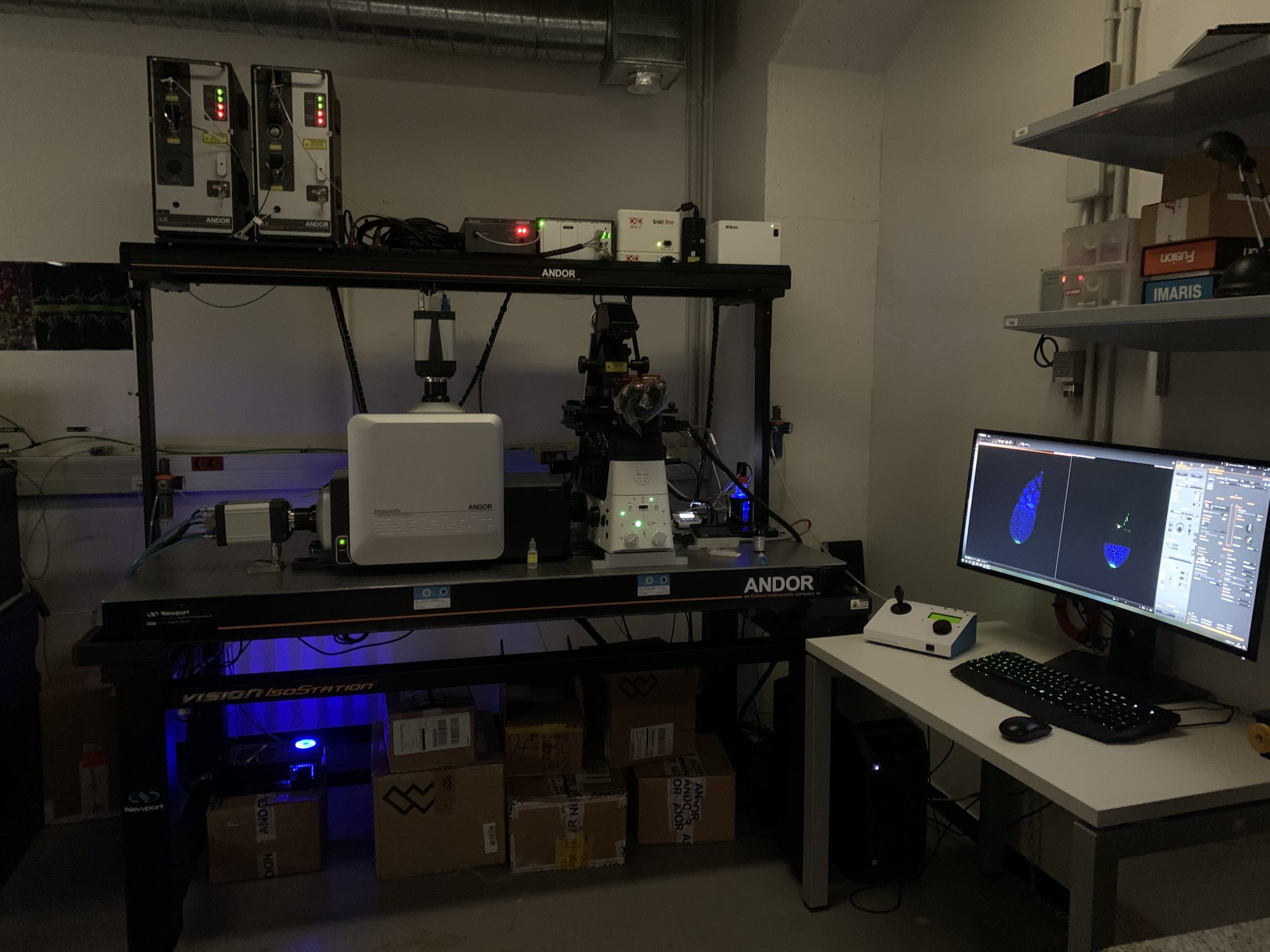
HIGH SPEED AND SUPER-RESOLUTION DRAGONFLY 505 (ANDOR)
Dragonfly 505 is a high speed & sensitivity multimodal confocal microscope whose applications range from live cell widefield and confocal imaging to single molecule localization, and from TIRF to whole embryo and thick tissue imaging. It delivers real time visualization for rapid specimen inspection, in parallel to GPU driven deconvolution to maximize resolution and throughput.

State subprogramme for Research Infrastructures and Scientific-Technical equipment Co-financed by the European Regional Development Fund (ERDF) with a grant of 614.919,65€
LSM780 CONFOCAL & MULTIPHOTON MICROSCOPE (CARL ZEISS)

The Zeiss Lsm780 is a confocal system with exceptional sensitivity. The GaAsP detector achieves 45% quantum efficiency compared to the 25% of conventional PMT detectors, and is capable of photon counting. The system´s illumination/detection design allows for simultaneous acquisition of up to ten dyes. The new optical design, the electronics concept and the user-friendly software make this confocal system fast, efficient and easy to use. In addition, the system is configured for Fluorescence Correlation Spectroscopy (FSC) and equipped with a MaiTai Deepsee IR laser for Multiphoton Microscopy (MPM).
Specifications
State subprogramme for Research Infrastructures and Scientific-Technical equipment Co-financed by the European Regional Development Fund (ERDF) with a grant of 614.919,65€
AF7000 MOTORIZED WIDEFIELD MICROSCOPE (LEICA MICROSYSTEMS)
The Leica AF6500 wide-field system offers a flexible platform for a wide range of conventional fluorescence applications, from routine imaging and documentation to high-speed live cell imaging. It is equipped with temperatura and CO2 control and uses the same easy-to-use software platform shared by the Leica confocal systems.

Specifications

Elena Rebollo

Tania García

PCB agreement *: Until December 2024 it guaranties free access to the Leica wide-field AF7000 microscope under the SCC self-service regime, introductions granted. The remaining equipment will be charged at the internal SCC-PCB price listed above.
General discounts: All users will benefit from a 50% discount during the overnight slots (11pm to 8am).
Non IBMB user´s may additionally include technical assistance (36.83 euros/hour).
Become a user
Registration
- Registration in Agendo is required to book the microscopes. Send an email to erabmc@ibmb.csic.es
Introductions and Permissions
- We provide Introductions to all the systems on a regular basis. Send a request to tgbmco@ibmb.csic.es
- The Nikon E600 microscope and the imaging workstations do not require permissions.
Access to the Booking Application
- Once you have booking permissions, you can access the Booking Application.
External Users
- External researchers should request an authorization to enter the PCB premises during non-working hours. Send a request to tgbmco@ibmb.csic.es
User guidelines
Here are some general guidelines we follow to make every day’s work smooth and efficient. If you need any special requirements, or have any useful suggestions, please contact us directly.
General
- Registration and Introduction to the systems by the Unit staff are compulsory for every user, either novice or advanced.
- If you cannot remember how to operate a system you were already introduced to, ask the stuff team for advice.
- Please do not enter in the facility any samples that are potentially hazardous or classified over S1 safety group.
- If you publish or present any work made or processed in any of our systems, please acknowledge the “Molecular Imaging Platform IBMB”..
Before you start
- Wipe off dirt and smear from the lens and the sample coverslip using lens tissue (soaked in ethanol:ether 2:1 if required).
- When environmental control is needed, make sure you have switched on temperature controller at least 30 min in advanced.
While working
- Check which immersion media are required for each lens you use, before you add any medium (oil, glycerol, water, air). Make sure corrections collars are properly set on the objective. If you don’t know how to do this, ask the facility stuff.
- Use moderate amounts (1 drop=50 µl) of immersion media. Immediately remove immersion media spilling over the objective rim.
- Clean immersion media from lenses with a dry lens tissue every time you change the sample. If you don’t do this, media (i.e. oil) will eventually overflow, causing damage to the objective and the microscope.
After you have finished
- Clean immersion media from every lens you used with a dry lens tissue.
- Clean immersion media form the stage holder.
- Always check up the reservations page to see whether somebody is booked after you.
- If you are last user of the day, switch off the equipment (fluorescence lamps, lasers, microscope controllers, cameras, softwares, PCs), according to the instructions provided by the facility stuff. Clean up the microscope bench from any residues (including dangerous glass coverslips and slides pieces). Do not leave your samples behind.
- Pay especial attention to the “system switch off” on Fridays; please make sure the lasers and fluorescence lamps are not left on over the weekend.
- Incidences should be reported to the unit stuff, either in person or by e-mail. Writing incidences in the logging sheet is not a guarantee of efficient assistance.
Booking Guidelines
- Booking of the microscope before use is mandatory, even if it is not being used by anybody else.
- Logging sheets (at the microscope) should be filled up and signed after your session.
- If you have to delete a booking, please do it at least 24 hours in advanced. Try to avoid short notice cancellations.
- Please send an e-mail to the system´s users list everytime you delete a booking. You can request us to be included in the cancellations notification lists of the different systems.
- If you delete the last booking of the day, ensure that the system is switched off (the previous user might not know that you are going to delete your booking).
- Try to set long time-lapse experiments during nights and weekends, especially if the system is saturated (remember to switch off devices not needed during time lapse like the HBO lamp on confocal systems).
- If you would like a long experiment to be saved by the next user, please contact him/her and get to an agreement.
- Indicate in the booking application whether you will use temperature control during your session, or any other requirement set on the reservations page.
Data transfer and storage
- When you save data locally, store it in a folder prepared to that purpose (normally it will be local hard disk D, folder “Users” or “Data”). Do not store data on the C: system hard disk, it may damage the system and reduce working memory.
- Transfer your data as soon as possible to avoid data loss, local hard disks will be automatically deleted in 15 days. Please remember that external memories/pendrives are not allowed due to the risk of viruses. Use the servers designated to that purpose.
Servers
Data will be automatically deleted in all MIP computers in 15 days, according to creation date. In order to transfer your files, use the servers below (no external memories are allowed)
“Caballa” PCB Server
Data will be stored in the server for 1 week, after that time they will be automatically deleted
How to get permissions
- Check if you are included at the PCB’s database (contact reception and ask, Helix: 934037172, recepcionhelix@pcb.ub.cat; Cluster: 934037171, recepcioncluster@pcb.ub.cat).
- Click on the following link: https://app.pcb.ub.es/usuaris/servlet/UsuariCrearFormulari12
- Create your new email address for PCB.
- In 2-3 days you will receive a letter by internal post with your username and temporary password (you can change it by clicking Here).
How to connect
Windows users
- Go to “My pc” and write in the address bar: \\caballa.adm.pcb.ub.es
- Click on the folder “microscopy”
- Insert your user name as: adm\username
- Type your password
Mac users
- Finder/Go/Connect to Server, write in the address bar:
smb://caballa.adm.pcb.ub.es - Insert your user name as: adm\username
- Type your password
- Click on the folder “microscopy”
Please disconnect from server once you have finished
Warning: do not click on “remember my password”, otherwise access will be blocked for other users !!!
MIP FTP Server
Only data acquired at MIP is allowed on this server. Users will need to delete their own data to manage their storage space
How to ask for credentials
- Send an email to tgbmco@ibmb.csic.es with cc: informatica@ibmb.csic.es
How to connect
Using Filezilla client (installed in all the facility computers).
- Open Filezilla and write the host (161.111.236.17), username and password, port: 22.
- Click “Quick connect”
- Before transferring your files, select the transfer default action “overwrite file”, to prevent files getting stuck in the queue
E. Rebollo publications list: https://bit.ly/2WbVguw
Computer Optimized Microscopy
Methods and Protocols
Automated Macro Approach to Quantify Synapse Density in 2D Confocal Images from Fixed Immunolabeled Neural Tissue Sections
Methods in Molecular Biology, vol. 2040, pp. 71-97
Automated Macro Approach to Remove Vitelline Membrane Autofluorescence in Drosophila Embryo 4D Movies
Methods in Molecular Biology, vol. 2040, pp. 71-97, pp 155-175
Open source code imaging contributions at https://github.com/MolecularImagingPlatformIBMB
MIP publications during 2019-2022
Mad3 modulates the G1 Cdk and acts as a timer in the Start network. Pérez AP, Artés MH, Moreno DF, Clotet J, Aldea M. Science Advances. 8(18) (May 2022)
The basement Membrane controls size and integrity of the Drosophila tracheal tubes. Klubmann-Fricke B-J; Martin-Bermudo MD* and Llimargas M*. Cell Reports, 39(4) (April 2022).
Pathway-specific effects of ADSL deficiency on neurodevelopment. Dutto I, Gerhards J, Herrera A, Souckova O, Škopová V, Smak JA, Junza A, Yanes O, Boeckx C, Burkhalter MD, Zikánová M, Pons S, Philipp M, Lüders J, Stracker TH. Elife (February 2022)
Gαq activation modulates autophagy by promoting mTORC1 signaling. Cabezudo S; Caballero A; Sanz Flores M, et al. Nature Communications (July 2021).
Coupled sterol synthesis and transport machineries at ER–endocytic contact sites. Encinar del Dedo J; Fernández-Golbano IM; Pastor L; Meler P; Rebollo E; Geli MI. Journal of Cell Biology 220(10) (July 2021).
EEF1A2 Controls Local Translation and Actin Dynamics in Structural Synaptic Plasticity. Mendoza MB, Gutierrez S, Ortiz R, Moreno DF, Dermit M, Dodel M, Rebollo E, Bosch M, Mardakheh FK, Gallego C. Science Signaling 14(691) (July 2021) .
Dbnl and β-catenin promote pro-N-cadherin processing to maintain apico-basal polarity. Herrera A, Menendez A, Torroba B, Ochoa A, Pons S. Journal of Cell Biology 220, no. 6 (June 2021): e202007055.
Cell intercalation driven by SMAD3 underlies secondary neural tube formation. Gonzalez-Gobartt E, Blanco-Ameijeiras J, Usieto S, Allio G, Benazeraf B, Martí E. Developmental Cell 56, no. 8 (April 2021):1147-1163.
AutoScanJ: A Suite of ImageJ Scripts for Intelligent Microscopy. Tosi, S; Stockinger, P; Godo, A; Rebollo, E; Bardia, L; Lladó, A; Colombelli, J. Frontiers in Bioinformatics (March 2021).
Stress Granules Display Bistable Dynamics Modulated by Cdk. Yahya, G, AP Pérez, MB Mendoza, E Parisi, DF Moreno, C Gallego, and M Aldea. Journal of Cell Biology 220, no. 3 (March 2021):e202005102.
Cholesterol alters mitophagy by impairing optineurin recruitment and lysosomal clearance in Alzheimer’s disease. Roca-Agujetas V, Barbero-Camps E, de Dios C, Podlesniy P, Abadin X, Morales A, Marí M, Trullàs R, Colell A. Molecular Neurodegeneration 16, no. 1 (March 2021): 15.
Development and Angiogenic Potential of Cell-Derived Microtissues Using Microcarrier-Template. Rubí-Sans G; Cano-Torres I; Pérez-Amodio S; Blanco-Fernández B; Mateos-Timoneda MA; Engel E. Biomedicines 9(3):232 (Feb 2021).
Condensin minimizes topoisomerase II-mediated entanglements of DNA in vivo. Dyson S, Segura J, Martínez-García B, Valdés A, Roca J. The EMBO Journal, 40, no. 1 (January 2021): e105393.
Multimerization of Zika Virus-NS5 Causes Ciliopathy and Forces Premature Neurogenesis. Saade, Murielle, Diego S. Ferrero, José Blanco-Ameijeiras, Elena Gonzalez-Gobartt, Marco Flores-Mendez, Victor M. Ruiz-Arroyo, Elena Martínez-Sáez, et al. Cell Stem Cell 27, no. 6 (December 3, 2020): 920-936.e8.
FoxO Maintains a Genuine Muscle Stem-Cell Quiescent State until Geriatric Age. García-Prat, Laura, Eusebio Perdiguero, Sonia Alonso-Martín, Stefania Dell’Orso, Srikanth Ravichandran, Stephen R. Brooks, Aster H. Juan, et al. Nature Cell Biology 22, no. 11 (November 2020): 1307–18.
Unravelling the Distinct Contribution of Cell Shape Changes and Cell Intercalation to Tissue Morphogenesis: The Case of the Drosophila Trachea. Casani, Sandra, Jordi Casanova, and Marta Llimargas. Open Biology 10, no. 11 (November 2020): 200329.
A Fraction of Barrier-to-Autointegration Factor (BAF) Associates with Centromeres and Controls Mitosis Progression. Torras-Llort, Mònica, Sònia Medina-Giró, Paula Escudero-Ferruz, Zoltan Lipinszki, Olga Moreno-Moreno, Zoltan Karman, Marcin R. Przewloka, and Fernando Azorín. Communications Biology 3, no. 1 (August 2020): 454.
A SMAD1/5-YAP signalling module drives radial glia self-amplification and growth of the developing cerebral cortex. Najas S, Pijuan I, Esteve-Codina A, Usieto S, Martinez JD, Zwijsen A, Arbonés ML, Martí E, Le Dréau G. Development 147, no. 13 (July 2020): dev187005.
Culture and Differentiation of Human Hair Follicle Dermal Papilla Cells in a Soft 3D Self-Assembling Peptide Scaffold. Betriu, Nausika, Claire Jarrosson-Moral, and Carlos E. Semino. Biomolecules 10, no. 5 (April 2020):684.
Mechanical Coordination Is Sufficient to Promote Tissue Replacement during Metamorphosis in Drosophila. Prat-Rojo, Carla, Philippe-Alexandre Pouille, Javier Buceta, and Enrique Martin-Blanco. The EMBO Journal 39, no. 3 (March 2020): e103594.
Proteostasis Collapse Halts G1 Progression and Delimits Replicative Lifespan. Moreno, David F., Kirsten Jenkins, Sandrine Morlot, Gilles Charvin, Attila Csikász-Nagy, and Martí Aldea. eLife 13, no .8 (September 2019): e48240.
PHF2 histone demethylase prevents DNA damage and genome instability by controling cell cycle progression of neural progenitors. Pappa S, Padilla N, Iacobucci S, Vicioso M, Álvarez de la Campa E, Navarro C, Marcos E, de la Cruz X, Martínez-Balbás MA. Proc Natl Acad Sci USA 116, no. 39 (September 2019): 19464-19473.
Anti-Siglec-1 Antibodies Block Ebola Viral Uptake and Decrease Cytoplasmic Viral Entry. Perez-Zsolt, Daniel, Itziar Erkizia, Maria Pino, Mónica García-Gallo, Maria Teresa Martin, Susana Benet, Jakub Chojnacki, et al. Nature Microbiology 4, no. 9 (September 2019): 1558–70.
Altered Cytoplasmic Maturation in Rescued in Vitro Matured Oocytes. Ferrer-Vaquer, A, M Barragán, A Rodríguez, and R Vassena. Human Reproduction 34, no. 6 (June 2019): 1095–1105.
Competition in the Chaperone-Client Network Subordinates Cell-Cycle Entry to Growth and Stress. Moreno, David F., Eva Parisi, Galal Yahya, Federico Vaggi, Attila Csikász-Nagy, and Martí Aldea. Life Science Alliance 2, no. 2 (April 2019): e201800277.
Dendritic Cells From the Cervical Mucosa Capture and Transfer HIV-1 via Siglec-1. Perez-Zsolt, Daniel, Jon Cantero-Pérez, Itziar Erkizia, Susana Benet, Maria Pino, Carla Serra-Peinado, Alba Hernández-Gallego, et al. Frontiers in Immunology 10 (April 2019): 825.
3D + Time Imaging and Image Reconstruction of Pectoral Fin During Zebrafish Embryogenesis. Nguyen, Hanh, Jaume Boix-Fabrés, Nadine Peyriéras, and Elena Kardash. Methods in Molecular Biology 2040 (March 2019): 135–153.
Automated Macro Approach to Quantify Synapse Density in 2D Confocal Images from Fixed Immunolabeled Neural Tissue Sections. Rebollo, Elena, Jaume Boix-Fabrés, and Maria L. Arbones. Methods in Molecular Biology 2040 (March 2019): 71–97.
Automated Macro Approach to Remove Vitelline Membrane Autofluorescence in Drosophila Embryo 4D Movies. Boix-Fabrés, Jaume, Katerina Karkali, Enrique Martín-Blanco, and Elena Rebollo. Methods in Molecular Biology 2040 (March 2019): 155–175.
Coincidence Analysis of Molecular Dynamics by Raster Image Correlation Spectroscopy. Moreno, David F., and Martí Aldea. Methods in Molecular Biology 2040 (March 2019): 375–384.
Impaired development of neocortical circuits contributes to the neurological alterations in DYRK1A haploinsufficiency syndrome. Arranz J; Balducci E; Arató K; et al; Arbonés ML. Neurobiology of Disease 127, pp.210-222 (2019). ISSN 0969-9961.
EQC2021-007200-P
Title: Supply and Installation of a High Speed Multiplexing FLIM Platform
Ministry of Sciences & Innovation (MICIN).
State Plan for Scientific and Technical Research and Innovation 2017-2020
Financed by the European Union: Recovery and resilience Funds.
PI: E. Rebollo
Ayuda EQC2021-007200-P financiada por:

EQC2018-004541-P
Supply and Installation of a High Speed and Super-resolution Imaging Platform
CPV code: 38515000-8
Ministry of Sciences, Innovation and Universities (MICIU)
StateSubprogramme for Research Infrastructures and Scientific-Technical equipment
Co-financed by the European Regional Development Fund (ERDF)
PI: E. Rebollo
CSIC13-4E-2065
Spectral confocal microscope for single molecule detection and 2-P nano-spectroscopy.
CPV code: 38515000-8
Ministry of Economy and Competitiveness (MINECO)
State Subprogramme for Research Infrastructures and Scientific-Technical equipment.
Co-financed by the European Regional Development Fund (FEDER)
PI: E. Rebollo
Collaboration Agreement with the Barcelona Sciences Parc.
This Platform has been set-up as a co-operation between the Molecular Biology Institute of Barcelona (IBMB) and the Barcelona Sciences Park (PCB), which ensures general services and maintenance for those systems covered by the PCB common services regime.