Oferta de contracte per realitzar la substitució del làser d'un microscopi LSM70/Axio Observer. Ha d'incloure…
INPhINIT 2018 J. Fernández Recio Position
Rational design of small-molecule inhibitors of amino acid transmembrane transporters for drug discovery
INPhINIT, the new PhD programme from “La Caixa” Foundation, opens today the call to recruit 57 early-stage researchers to the top Spanish centres and units of research. The Structural Biology Unit, as a María de Maeztu unit of excellence, offers four PhD positions in structural biology cutting-edge research projects.
The aim of this programme is to support the talented young researchers and to foster the innovative and high-quality Spanish research. The researchers will enjoy a 3-year contract in a research centre among those awarded by the Spanish Ministry of Economy and Competitiveness (“Severo Ochoa” centres of excellence and “Maria de Maeztu” units of excellence) and the Spanish Ministry of Health (“Carlos III centres of excellence”) that offer a competitive environment to develop a research of excellence.
Research project/ Research Group description
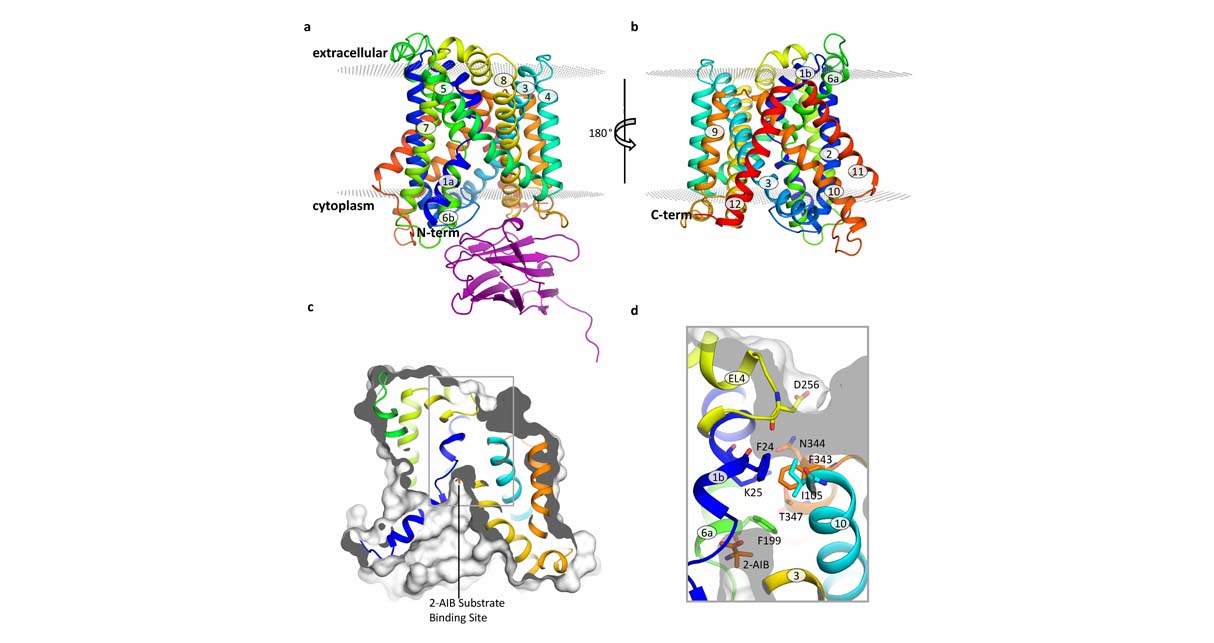
Heteromeric Amino-acid Transporters (HAT) constitute a family of transmembrane proteins that are involved in serious human pathologies, like amino acidurias, cancer, tumor cell growth, herpesvirus infection or cocaine addiction relapse, and thus are attractive targets for therapeutic drug discovery. For instance, inhibition of xCT (glutamate/cystine exchanger) has been proposed as target for human gliomas, which have a dismal prognosis with very few treatment options. The proposed project will focus on the discovery of specific inhibitors for several members of the HAT family. The information acquired for each member will be useful to understand the inhibition of the other members. In the case of xCT, the group started a structure-based drug discovery program within a collaborative project funded by Fundació La Marató de TV3. Preliminary results based on computational modeling and cell biology analysis have found several small compounds capable of inhibiting xCT and chemically different to previously known inhibitors (unpublished).
The research group “Interactions and Protein Docking” was created in 2004 by Juan Fernández Recio. The group recently moved to the Maria de Maeztu Structural Biology Unit at the Barcelona Molecular Biology Institute (IBMB) when the PI joined the CSIC. The research of the group focuses on the theoretical study of biological phenomena at molecular level and the development of computer tools for the analysis, structural prediction and characterization of protein interactions and protein-protein binding sites. The research team is actively involved in many multi-disciplinary collaborative projects for the study of systems of biomedical interest. The computer tools developed in the group are free for academics and/or available through different web services and databases, and they consistently rank highly in the CAPRI experiment, an ongoing series of blind tests that evaluate the research community ability to model protein-protein complexes.
Job position description
The main objective of the job is to optimize xCT inhibitors that were previously identified in the group, discover new inhibitors for other HAT proteins, and use all this information as a basis to design new drug candidates against different pathologies, such as human glioma. This will be achieved by computational structural modelling of HAT proteins and their interactions. It will be important to consider the different conformational states of HAT proteins (inward- and outward-facing, etc.), for which there are several structural templates. In addition to the goal of obtaining drug candidates, the resulting models and proposed analyses in this project will help to understand the mechanism of amino acid transmembrane transport through this system.
This will be part of a fruitful collaboration with Prof. Manuel Palacín (Institute for Research in Biomedicine of Barcelona), which in the past has involved several granted projects (Fundació La Marató de TV3, agreement with SIDRA Medical and Research Center from Qatar). The candidate will work in a highly interdisciplinary environment. He/she will learn and apply state-of-the-art structural biology and computational techniques (homology modeling, protein docking, molecular dynamics, virtual screening, QSAR), which will need to be integrated with cell biology experiments (transport activity and cell viability, chemical synthesis, pharmacodynamics studies and cheminformatics analyses). This will need constant interaction with wet laboratory researchers, interactive feed-back from experiments, and open mind to bring computational tools into practical applications.
For more information about the call, visit INPhINIT website
Deadline: 01/02/2018
This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Sklowdowska-Curie grant agreement No. 713673